Raw data in. Clean results out.
Upload your qPCR file, get publication-ready analysis in seconds. No installs, no spreadsheets, no headaches.
How It Works
From raw file to clean results in three steps.
Upload
Drop your .xlsx, .xls, .csv, or native instrument file. We auto-detect the format.
Configure
Pick your reference gene, control group, and analysis method. Or let the defaults run.
Voila
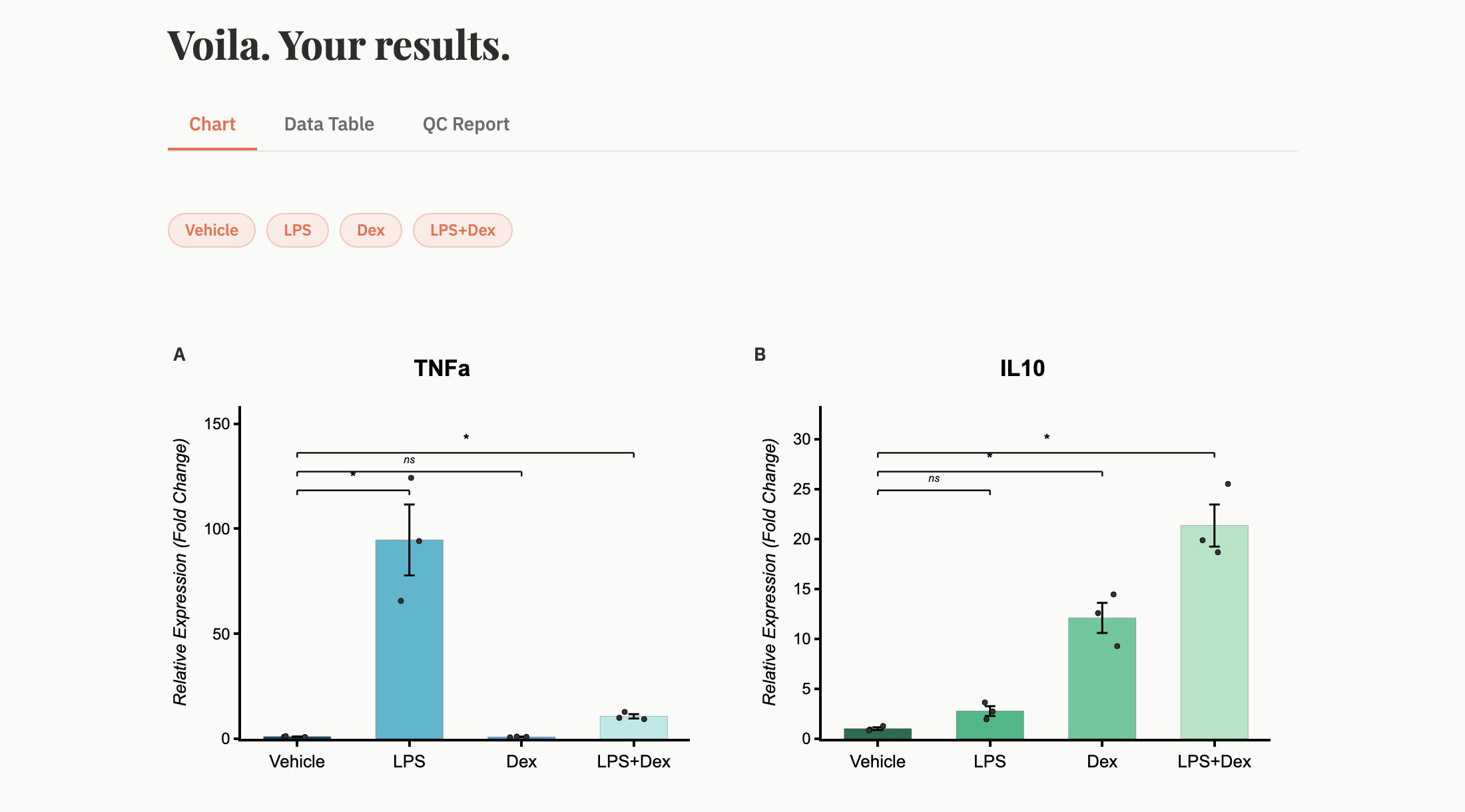
Get fold-change tables, QC reports, and publication-ready figures. Export or share instantly.
No sign-up required. Your data never leaves your browser.
Try it now
Upload your file and see results in seconds.
Drop your spreadsheet here
or click to browse
.xlsx, .xls, or .csv — QuantStudio, CFX, LightCycler, ABI 7500, Biomark, and more
Built for bench scientists
Everything you need for rigorous, reproducible qPCR analysis.
Automated QC Report
Eight diagnostic checks run instantly: NTC contamination, replicate scatter, reference-gene stability, late Ct, undetermined wells, extreme fold-change, amplification efficiency, and identical-Ct flags.
Publication Figures
Export bar charts styled for Nature, Cell, or PNAS with one click. Customize colors, fonts, error bars, and significance markers, then download as SVG or PNG.
Single & Multi-Plate
Analyze one plate or combine biological replicates across multiple plates in a single run. Mix instrument formats freely — VoilaPCR merges and normalizes automatically.
Supported Instruments
Auto-detected on upload. No manual format selection needed.
QuantStudio 3/5/6/7
Thermo Fisher
StepOne / StepOnePlus
Applied Biosystems
ABI 7500 / 7500 Fast
Applied Biosystems
CFX96 / CFX384 / Opus
Bio-Rad
LightCycler 96 / 480
Roche
Rotor-Gene Q / 6000
QIAGEN
Stratagene Mx3000P/3005P
Agilent
Mastercycler realplex
Eppendorf
Mic qPCR (48-well)
Bio Molecular Systems
BioMark HD / EP1
Fluidigm
Generic CSV
Any instrument
Built for trust
Your data never leaves your browser
All parsing, QC, and analysis run entirely in JavaScript on your machine. No file is uploaded to any server. Your raw data is never transmitted, stored, or accessible to anyone but you.
MIQE-aware diagnostics
Our QC checks are aligned with the MIQE guidelines (Bustin et al., 2009). Each diagnostic references the relevant MIQE criterion so reviewers can verify compliance.
Cite VoilaPCR
Use this citation in your methods section: